All content on this site is intended for healthcare professionals only. By acknowledging this message and accessing the information on this website you are confirming that you are a healthcare professional. If you are a patient or carer, please visit the Lymphoma Coalition.
The Lymphoma Hub website uses a third-party service provided by Google that dynamically translates web content. Translations are machine generated, so may not be an exact or complete translation, and the Lymphoma Hub cannot guarantee the accuracy of translated content. The Lymphoma Hub and its employees will not be liable for any direct, indirect, or consequential damages (even if foreseeable) resulting from use of the Google Translate feature. For further support with Google Translate, visit Google Translate Help.
The Lymphoma & CLL Hub is an independent medical education platform, sponsored by AbbVie, BeOne Medicines, Miltenyi Biomedicine, Nurix Therapeutics, Roche, Sobi, and Thermo Fisher Scientific and supported through educational grants from Bristol Myers Squibb, Eli Lilly, and Pfizer. Funders are allowed no direct influence on our content. The levels of sponsorship listed are reflective of the amount of funding given. View funders.
Now you can support HCPs in making informed decisions for their patients
Your contribution helps us continuously deliver expertly curated content to HCPs worldwide. You will also have the opportunity to make a content suggestion for consideration and receive updates on the impact contributions are making to our content.
Find out more
Create an account and access these new features:
Bookmark content to read later
Select your specific areas of interest
View lymphoma & CLL content recommended for you
AACR 2017 | Poster 3939/6 – two evolutionary models proposed for development of drug resistance between primary and relapsed Diffuse Large B-Cell Lymphoma
At the American Association for Cancer Research (AACR) annual meeting in Washington, DC, USA, on Tuesday 4th April, a poster session titled “Tumor Evolution and Heterogeneity 2” took place.
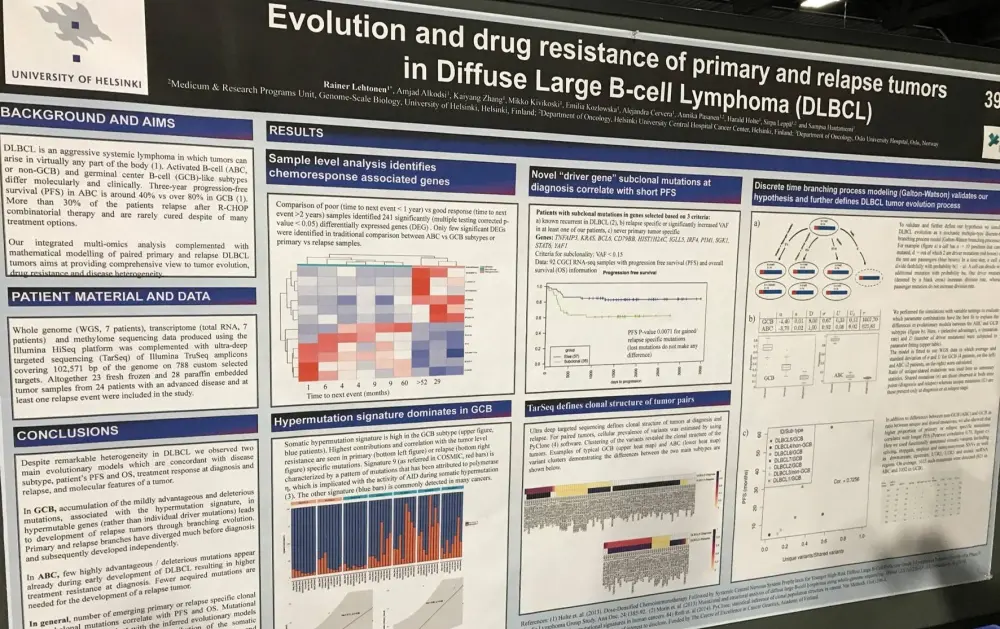
One of the posters on display (3939 / 6) was titled “Evolution and drug resistance of primary and relapse tumors in Diffuse Large B-Cell Lymphoma” by Rainer Lehtonen from the University of Helsinki, Helsinki, Finland, and colleagues.
This group carried out integrated multi-omics (whole genome, transcriptome, and methylome) analyses complemented with mathematical modelling of pared primary and relapsed DLBCL tumors. They aimed to provide a comprehensive view of tumor evolution, drug resistance, and disease heterogeneity. In total, 23 fresh frozen and 28 paraffin embedded samples from 24 advanced disease patients with at least one relapse were included.
Key Highlights:
- When comparing patients with time to next event <1yr versus >2yrs, 241 significantly (P < 0.05) differentially expressed genes (DEGs)
- Divergent evolution prominent in GCB whereas subclonal mutations are rare in ABC subtype
- Few genomic differences detected between primary and relapse tumors (P = 0.047, Pearson's Chi-squared test on 7 patients with WGS data); suggests pre-treatment drug resistance is likely caused by global hypo- or hypermethylation at relapse stage
- In GCB subtype, primary and relapse branches largely diverge before diagnosis and subsequently develop independently
- Number of emerging primary or relapse specific clonal and subclonal mutations correlate with PSF and OS
- Contribution of the somatic hypermutation signature was higher in resistant tumors and correlated with time to disease progression
- Identified 72 relapse tumor related genes including many known B-Cell Lymphoma genes such as CD79B, IGLL5, PIM1, SGK1
- Patients with subclonal mutations in these genes had shorter PSF and OS (P = 0122 and 0.0071, respectively)
The group concluded the poster by hypothesizing two evolutionary models, which take into account disease subtype, PFS and OS, treatment response at diagnosis and relapse, and molecular features:
- GCB: Buildup of mildly advantageous and deleterious mutations in hypermutable genes, associated with hypermutation signature, and not in individual driver mutations. This results in development of relapsed tumors through branching evolution. Divergence of primary and relapse branches thought to occur long before diagnosis and then develop independently
- ABC: A small number of highly advantageous/deleterious mutations occur early in DLBCL development resulting in higher resistance to treatment at diagnosis. Less acquired mutations are needed for relapsed tumor to develop
References
Please indicate your level of agreement with the following statements:
The content was clear and easy to understand
The content addressed the learning objectives
The content was relevant to my practice
I will change my clinical practice as a result of this content
Your opinion matters
In your experience, what is the average vein-to-vein time when treating patients with DLBCL with a reimbursed CAR T-cell therapy (from apheresis to infusion)?

