All content on this site is intended for healthcare professionals only. By acknowledging this message and accessing the information on this website you are confirming that you are a Healthcare Professional. If you are a patient or carer, please visit the Lymphoma Coalition.
The lym Hub website uses a third-party service provided by Google that dynamically translates web content. Translations are machine generated, so may not be an exact or complete translation, and the lym Hub cannot guarantee the accuracy of translated content. The lym and its employees will not be liable for any direct, indirect, or consequential damages (even if foreseeable) resulting from use of the Google Translate feature. For further support with Google Translate, visit Google Translate Help.
The Lymphoma & CLL Hub is an independent medical education platform, sponsored by AbbVie, BeOne Medicines, Miltenyi Biomedicine, Nurix Therapeutics, Roche, Sobi, and Thermo Fisher Scientific and supported through educational grants from Bristol Myers Squibb, Lilly, and Pfizer. Funders are allowed no direct influence on our content. The levels of sponsorship listed are reflective of the amount of funding given. View funders.
Now you can support HCPs in making informed decisions for their patients
Your contribution helps us continuously deliver expertly curated content to HCPs worldwide. You will also have the opportunity to make a content suggestion for consideration and receive updates on the impact contributions are making to our content.
Find out more
Create an account and access these new features:
Bookmark content to read later
Select your specific areas of interest
View lymphoma & CLL content recommended for you
iwCLL 2017 | Ongoing IGHV-D-J mutations result in substantial clonal complexity and presents as a measure of genomic instability in CLL
On 14th May 2017, the “Role of the B-Cell Receptor and Other Signaling Pathways in CLL” took place at iwCLL, and was co-chaired by Nicholas Chiorazzi (The Feinstein Institute for Medical Research, Manhasset, NY, USA) and Kostas Stamatopoulos (Center for Research and Technology Hellas, Thessaloniki, Greece).
“Intraclonal Diversification and Evolution in Chronic Lymphocytic Leukemia Patients by High-Throughput Sequencing of IGHV-D-J Rearrangements” was a talk presented during this session by Davide Bagnara from the University of Genova, Italy.
It has been previously reported in Sanger sequencing studies than IGHV-D-J of CLL clones can undergo intra-clonal heterogeneity.
This group performed high-throughput sequencing on the IGHV-D-J repertoire of FACS sorted CD19+CD5+ cells from 39 patients with treatment naïve CLL. From mRNA, and using a set of primers covering every IGHV gene, full-length IGHV-D-J repertoire was amplified. Unique Molecular Identifiers (for error) and Polymerase Chain Reaction (for correction) were used to prepare libraries. The CD19+CD5+ compartment in patients with CLL contains both leukemic and non-leukemia B-cell clones.
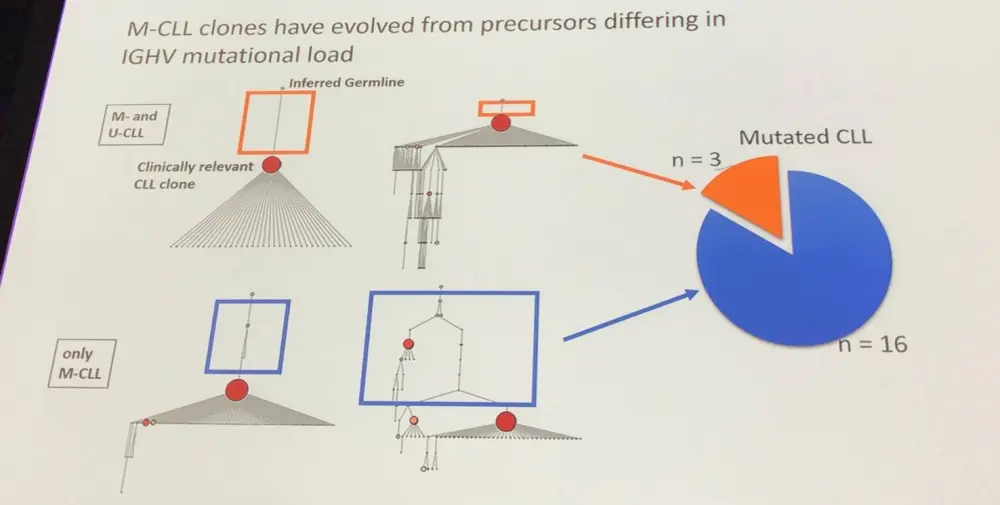
- CLL clones in the ex vivo environment without stimulation displayed intraclonal diversity of leukemic IGHV-D-J rearrangement, indicating ongoing somatic mutations
- This was observed in Unmutated IGHV CLL (U-CLL) and Mutated IGHV CLL (M-CLL) clones
- In a group of M-CLL cases, it appeared that the CLL clone evolved from a precursor differing in IGHV mutational status
- Two groups identified:
- Almost all subclones derive directly from the CLL clone
- A number of subclones develop in a complex, branching, phylogenetic tree; not directly deriving from one CLL clone
- When analyzing the change in proportion of a clone at two different time points:
- An increase in CLL clone indicates it has gained an advantage over the other subclones, not present in a previous state
- A decrease in CLL clone indicated one or more subclones numerically competed with it, indicating ongoing clonal diversification
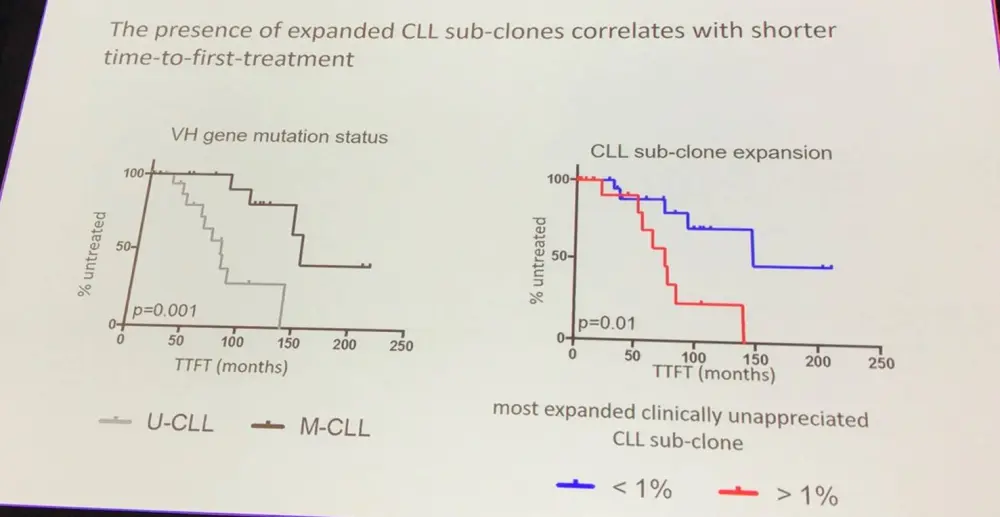
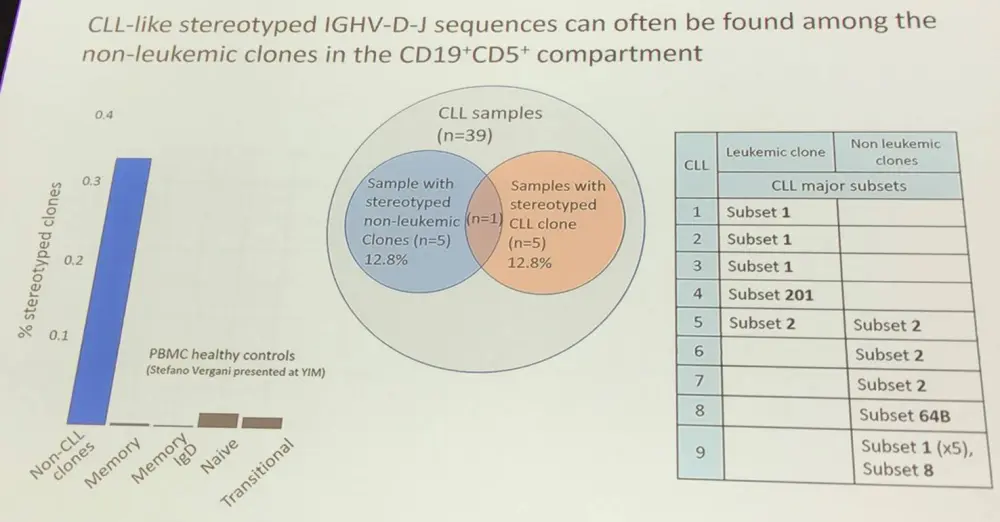
- It was found that CD19+CD5+ non-leukemic B-cells could be oligoclonally expanded
- CLL presents stereotypical IGHV-D-J sequences that can often be observed in clones of the non-leukemia CD19+CD5+ compartment

In conclusion, CLL clones diversify in vivo, acquiring IGHV-D-J mutations resulting in a level of clonal complexity not fully comprehended thus far, and takes place in both U- and M-CLL. The group hypothesized that the ongoing evolution of IGHV-D-J is likely due to Activation Induced Deaminase. Moreover, they suggested that ongoing IGHV-D-J could be used as a marker of DNA changes taking place across the genome and so presents as a measure of genomic instability.


References
Please indicate your level of agreement with the following statements:
The content was clear and easy to understand
The content addressed the learning objectives
The content was relevant to my practice
I will change my clinical practice as a result of this content

