All content on this site is intended for healthcare professionals only. By acknowledging this message and accessing the information on this website you are confirming that you are a Healthcare Professional. If you are a patient or carer, please visit the Lymphoma Coalition.
The lym Hub website uses a third-party service provided by Google that dynamically translates web content. Translations are machine generated, so may not be an exact or complete translation, and the lym Hub cannot guarantee the accuracy of translated content. The lym and its employees will not be liable for any direct, indirect, or consequential damages (even if foreseeable) resulting from use of the Google Translate feature. For further support with Google Translate, visit Google Translate Help.
The Lymphoma & CLL Hub is an independent medical education platform, sponsored by AbbVie, BeOne Medicines, Miltenyi Biomedicine, Nurix Therapeutics, Roche, Sobi, and Thermo Fisher Scientific and supported through educational grants from Bristol Myers Squibb, Lilly, and Pfizer. Funders are allowed no direct influence on our content. The levels of sponsorship listed are reflective of the amount of funding given. View funders.
Now you can support HCPs in making informed decisions for their patients
Your contribution helps us continuously deliver expertly curated content to HCPs worldwide. You will also have the opportunity to make a content suggestion for consideration and receive updates on the impact contributions are making to our content.
Find out more
Create an account and access these new features:
Bookmark content to read later
Select your specific areas of interest
View lymphoma & CLL content recommended for you
iwCLL 2017 | Recurrent somatic mutations and their functions in CLL
At iwCLL 2017, on Monday 13th May, the second half of the session titled “Unraveling the factors leading to the development of CLL”, and was chaired by Emili Montserrat (Hospital Clinic of Barcelona) and John Gribben (Barts Cancer Institute).
Catherine Wu, from the Dana-Farber Cancer Institute, gave a talk during this session titled: “Functional Evaluation of Key Recurrent Somatic Mutations in CLL.”
The talk began by outlining that CLL follows a clonal evolution:
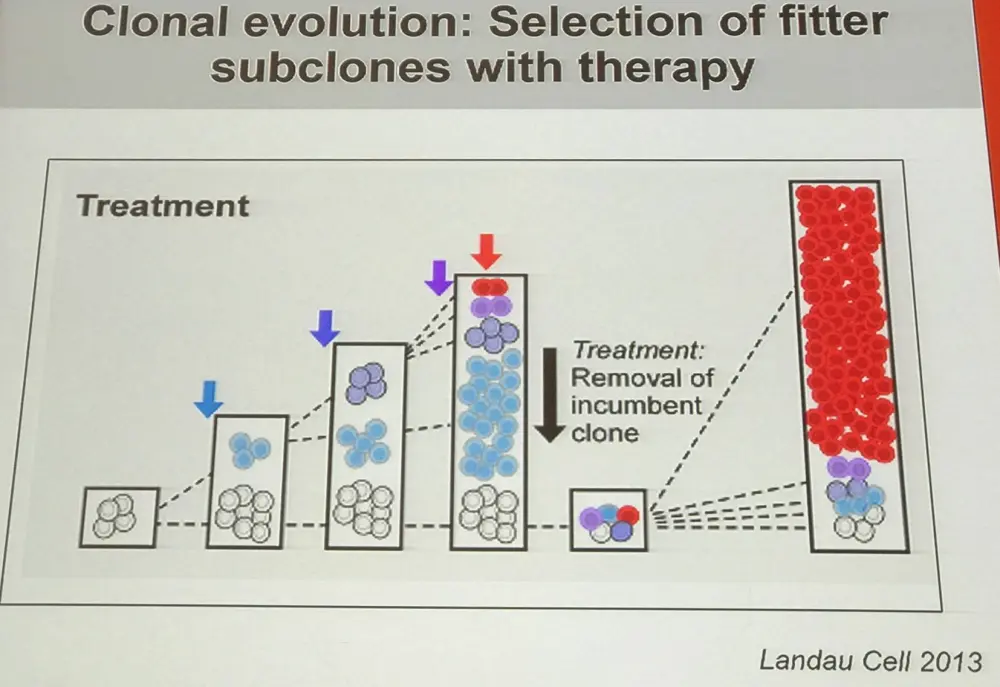
How are specific somatic mutations contributing to evolution and cellular fitness?
Firstly, SF3B1 mutation independently predicts poor prognosis (Landau et al. 2015). Mutation of SF3B1 (the catalytic core of the spliceosome) efficiently and subtly alters splicing and gene expression in a range of pathways.
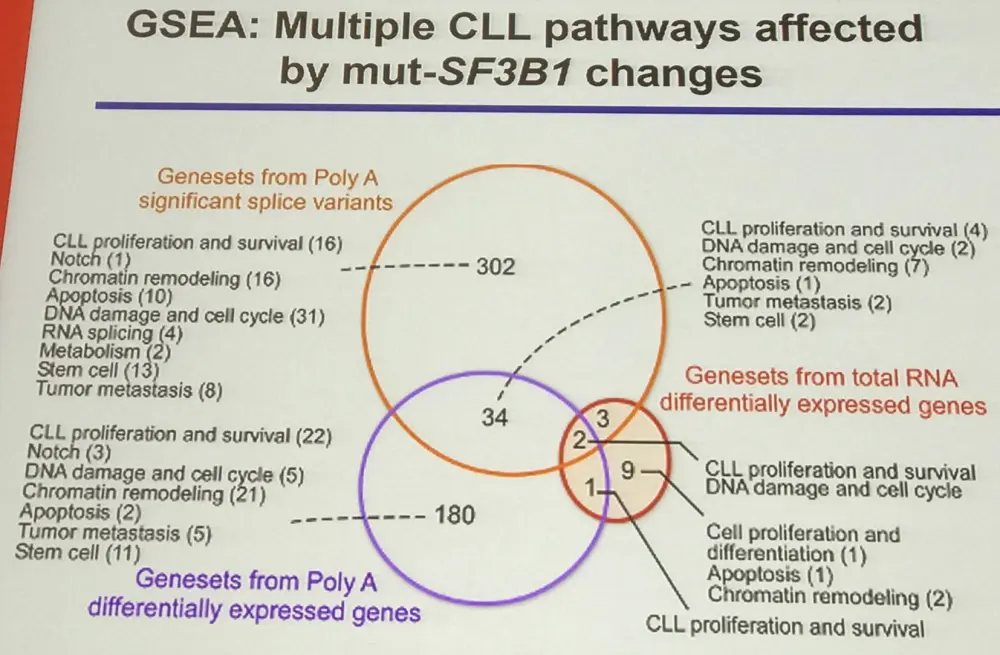

Surprisingly, the Notch signaling pathway has been found to be activated with mutant SF3B1 as well as subtle changes in DNA damage response and telomerase activity, but not in RNA splicing function, Wnt pathway, cell cycle, or apoptosis (Wang et al. 2016).
TERT, TERC, KLF8, and Alt-DVL2 are examples of gene expression and splice variants that impact on CLL pathways. In-frame deletion in DVL2 (disheveled; a key mediator of Wnt signaling) is associated with SF3B1 mutation and can inhibit Notch signaling in vivo by inhibiting CSL (CBF1, Suppressor of Hairless, LAG-1) transcription factors downstream of Notch receptors (Collu et al. 2012).
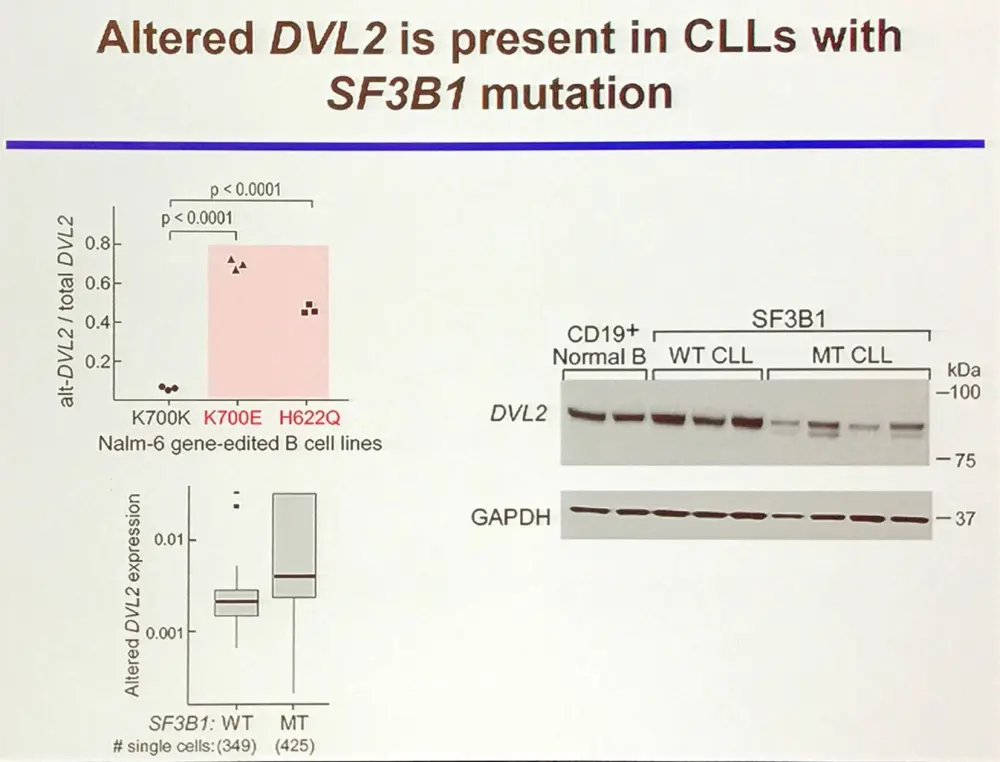
Mutations in SF3B1 is predominantly a subclonal event, and likely impacts on cellular fitness and the evolutionary capacity of the CLL cell. In CLL, this requires co-operation with additional mutations.
How do somatic mutations contribute to natural progression?
Uncontrolled growth is one of the key hallmarks of cancer; however, the rate and pattern of growth can differ between different types of cancer. A group, part of the CLL Research Consortium, collected serial samples (average = 4) from 15 patients from time of diagnosis to first therapy. Overall, 38 macroscopic subclones were identified.
In summary, comprehensive genetic characterization of CLL demonstrates high genetic heterogeneity amongst individuals. Moreover, future functional evaluation of somatic mutations should reveal the critical oncogenic checkpoints in CLL.
References
Please indicate your level of agreement with the following statements:
The content was clear and easy to understand
The content addressed the learning objectives
The content was relevant to my practice
I will change my clinical practice as a result of this content

