All content on this site is intended for healthcare professionals only. By acknowledging this message and accessing the information on this website you are confirming that you are a healthcare professional. If you are a patient or carer, please visit the Lymphoma Coalition.
The Lymphoma Hub website uses a third-party service provided by Google that dynamically translates web content. Translations are machine generated, so may not be an exact or complete translation, and the Lymphoma Hub cannot guarantee the accuracy of translated content. The Lymphoma Hub and its employees will not be liable for any direct, indirect, or consequential damages (even if foreseeable) resulting from use of the Google Translate feature. For further support with Google Translate, visit Google Translate Help.
The Lymphoma & CLL Hub is an independent medical education platform, sponsored by AbbVie, BeOne Medicines, Miltenyi Biomedicine, Nurix Therapeutics, Roche, Sobi, and Thermo Fisher Scientific and supported through educational grants from Bristol Myers Squibb, Eli Lilly, and Pfizer. Funders are allowed no direct influence on our content. The levels of sponsorship listed are reflective of the amount of funding given. View funders.
Now you can support HCPs in making informed decisions for their patients
Your contribution helps us continuously deliver expertly curated content to HCPs worldwide. You will also have the opportunity to make a content suggestion for consideration and receive updates on the impact contributions are making to our content.
Find out more
Create an account and access these new features:
Bookmark content to read later
Select your specific areas of interest
View lymphoma & CLL content recommended for you
The role of genetic drivers of DLBCL in the tumor microenvironment
Several factors influence the tumor microenvironment (TME) in diffuse large B-cell lymphoma (DLBCL), including genetic factors in lymphoma cells.1 Understanding how these genetic factors influence the TME could identify novel therapeutic targets, and improve the efficacy of immunotherapies, such as chimeric antigen receptor T-cell therapies.1
Here, we summarize a review published by Cerchietti.1 in Blood on the role of genetic drivers of DLBCL in the lymphoma microenvironment (LME).
Classification of LME
Using artificial intelligence-based computational analysis of transcriptional signatures, researchers have identified four major categories of LMEs in DLBCL with distinct characteristics (Figure 1):
- Germinal center-like LME (GC-LME)
- Mesenchymal LME (MS-LME)
- Inflamed and immunosuppressive LME (IN-LME)
- Depleted LME (DP-LME)
Figure 1. DLBCL LME categories*
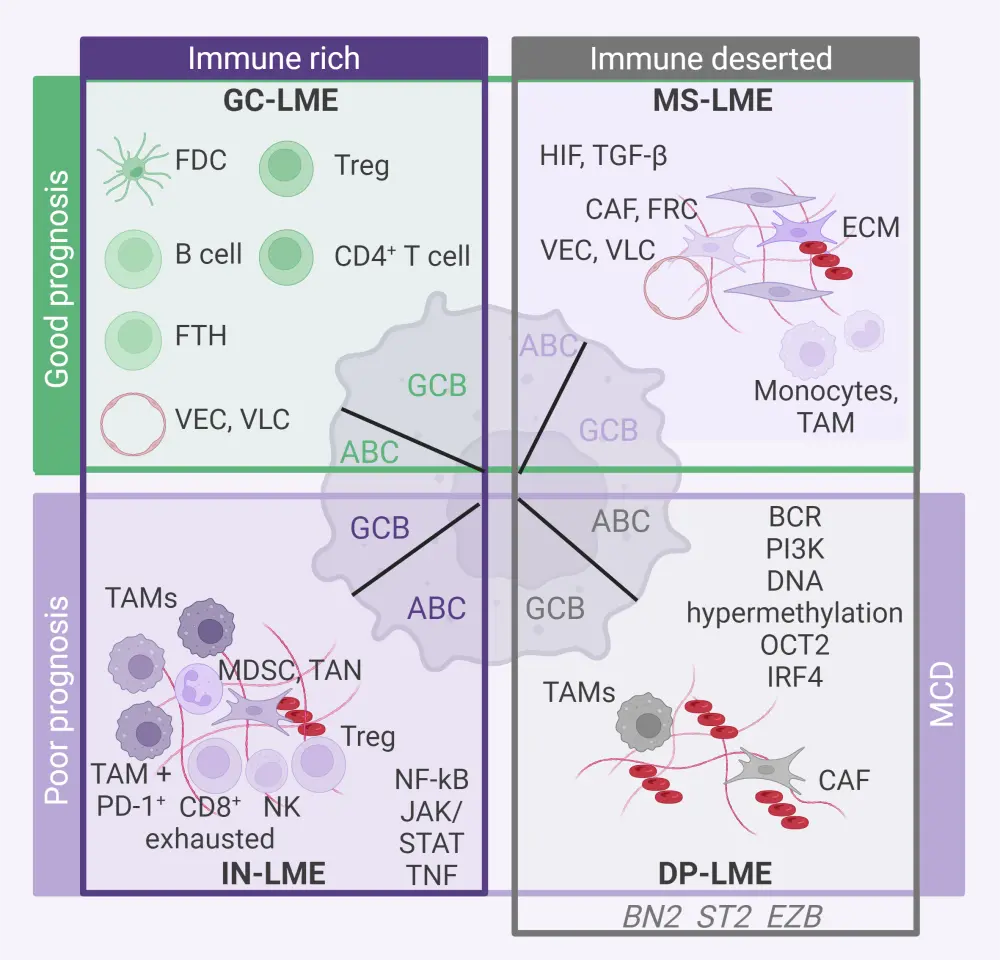
ABC, activated B-cell; BCR, B-cell receptor; CAF, cancer-associated fibroblast; DLBCL, diffuse large B-cell lymphoma; DP, depleted; ECM, extracellular matrix; FDC: follicular dendritic cell; FTH, follicular T-helper cell; FRC, fibroblastic reticular cell; GC, germinal center-like; GCB, germinal center B-cell; IN, inflammatory and immunosuppressive; IRF4, interferon regulatory factor 4; JAK, Janus kinase; LME, lymphoma microenvironment; MDSC, myeloid derived suppressor cell; MS, mesenchymal; NF-Κb, nuclear factor kappa B; NK, natural killer cell; OCT2, organic cation transporter 2; PD-1, programmed cell death-1; PI3K, phosphoinositide 3 kinase; STAT, signal transducer and activator of transcription; TAM, tumor-associated macrophage; TAN, tumor-associated neutrophil; TGF-β, transforming growth factor-β; TNF, tumor necrosis factor; Treg, regulatory T cell; VEC, vascular endothelial cell; VLC, vascular lymphatic cell.
*Adapted from Cerchietti.1
Impact of mutations on the TME
Mutations can affect the TME through several mechanisms, including:
- direct mutations of membrane ligands or receptors;
- mutations affecting signaling pathway transducers or scaffolding molecules; and
- expression or secretion of short- and long-range acting molecules.
The majority of genetic changes affecting the TME have a combination of direct and indirect, and short- and long-range effects on the TME, and TME changes are often influenced by multiple genetic alterations.
Direct effects on the TME
- Disruptions of antitumor immune mechanisms play a significant role in the transformation of the TME.
- The majority target the antigen processing and presentation pathway, and costimulatory and coinhibitory molecules.
- The most common antigen processing and presentation pathway alteration in DLBCL is the lack of expression of the class I major histocompatibility complex (roughly 40–75% of patients).
Indirect effects on the TME
- Mutations that affect the transcriptional regulators (e.g., CIITA) or epigenetic regulators (e.g., EZH2, CREBBP, KMT2D, EP300) can alter the expression of ligands and receptors.
Mutations that modify the secretome and metabolism of cancer cells
- Mutations that result in the release of short- and long-range acting molecules into the extracellular space can alter the TME.
- TP53 loss of heterozygosity and loss of function mutations can result in the modification of cellular membrane proteins and the secretome of cancer cells.
Impact of genotypic variants and alterations in non-tumor cells on the TME
- The genotypic variants and functional alterations in non-tumor cells can impact their susceptibility to reprogramming by cancer cells.
- Age and ethnicity can be associated with differences in the TME, possibly due to functional alterations in the genome of aged cells and the presence of single nucleotide variants.
Crosstalk of cancer and LME cells during progression
- While cancer cells can alter the TME during progression, the TME conditions can also result in changes in the cancer cells. This interaction between cancer and LME cells leads to crosstalk, causing the cells to evolve in a process of co-selection.
| Key learnings |
|
References
Please indicate your level of agreement with the following statements:
The content was clear and easy to understand
The content addressed the learning objectives
The content was relevant to my practice
I will change my clinical practice as a result of this content
Your opinion matters
What is the primary reason you use bridging therapy in patients with DLBCL awaiting CAR T-cell therapy?

